
| Microexon ID | Pt_14:5389241-5389255:+ |
| Species | Populus Trichocarpa | Coordinates | 14:5389241..5389255 |
| Microexon Cluster ID | MEP42 |
| Size | 15 |
| Phase | 0 |
| Pfam Domain Motif | bHLH-MYC_N |
| Structure of Microexon-tag (flanking exon, microexon, flanking exon sizes) | 48,15,45 |
| Microexon location in the Microexon-tag | 2 |
| Microexon-tag DNA Seq | GCWMAYKATGCMGAYAGYAAARTYTTYTCTCGMKCTHTKCTWGCMAAGAGTGCWTSWATTCAGACWGTGGTRTGYWTTCCYYWYMTRGRYGGYGTBRTTGARCTWGGY |
| Logo of Microexon-tag DNA Seq | 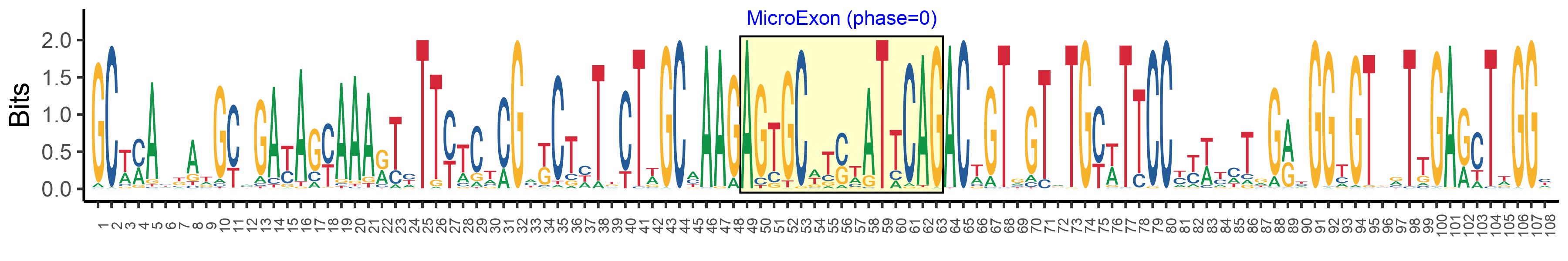 |
| Alignment of exons | 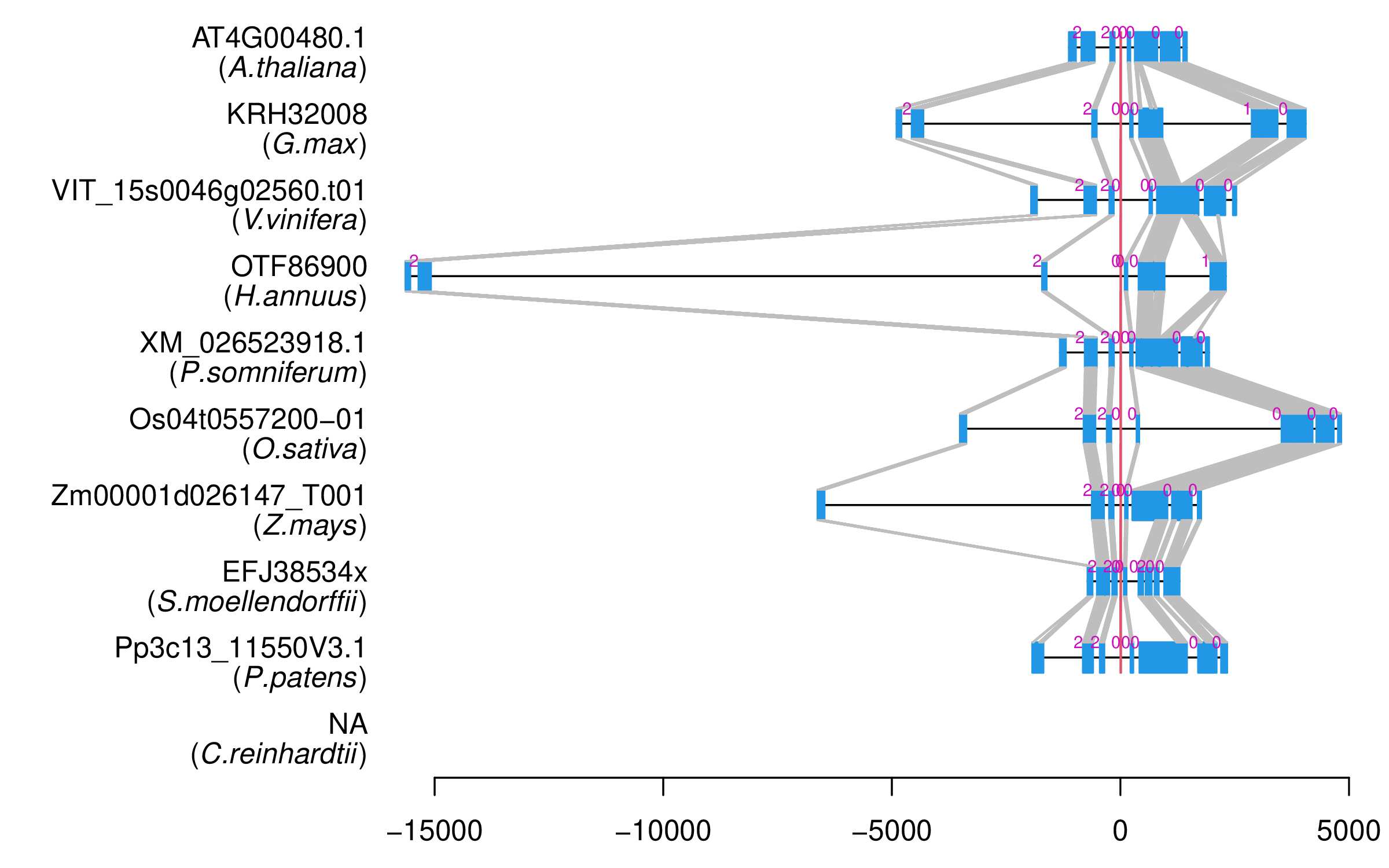 |
| Microexon DNA seq | AGCGCATCAATTCAG |
| Microexon Amino Acid seq | SASIQ |
| Microexon-tag DNA Seq | GCTCAATATGCAGATAGCAAAGTATTCTCTCGCTCTTTACTCGCAAAGAGCGCATCAATTCAGACTGTGGTGTGTTTTCCCTATCTTGAGGGAGTAATAGAGCTAGGT |
| Microexon-tag Amino Acid Seq | AQYADSKVFSRSLLAKSASIQTVVCFPYLEGVIELG |
| Microexon-tag spanning region | 5389046-5389792 |
| Microexon-tag prediction score | 0.9573 |
| Overlapped with the annotated transcript (%) | 100 |
| New Transcript ID | Potri.014G083900.2.v4.1x |
| Reference Transcript ID | Potri.014G083900.2.v4.1 |
| Gene ID | Potri.014G083900.v4.1 |
| Gene Name | NA |
| Transcript ID | Potri.014G083900.2.v4.1 |
| Protein ID | |
| Gene ID | Potri.014G083900.v4.1 |
| Gene Name | |
| Pfam domain motif | bHLH-MYC_N |
| Motif E-value | 4.70E-54 |
| Motif start | 15 |
| Motif end | 195 |
| Protein seq | >Potri.014G083900.2.v4.1 MAPVVQGQQVVPDNLRKQLAVAVRSVQWSYAVFWSQSTRQQGVLEWGDGYYNGDIKTRKVEAMELKADKIGLQRSEQLRE LYESLLEGETGLQATRSSPALSPEDLSDEEWYYLVCMSFVFNPGEGLPGRALANKQPIWLCNAQYADSKVFSRSLLAKSA SIQTVVCFPYLEGVIELGVTELVTEDPGLIQHIKASLLDFSKPVCSDKSFSAAHNKDDDKDPMSTRISHEIVDTLALENL YTPTEDIESEQEGINYLHGNVCEEFNRNSPDDFSNGYEHNLVTEDSFMLEDLKEGASQVQSWHSMDDEFSDDVRDSMNSS DCISEVFVKQGKVVPSSKGKDISHLQLKVLQEGNHTKLSSLDPGADDDLHYRRTAFVILKSSSQLIENPCFQSGDYKSSF VGWKKGAADGYKPRIQQKMLKKILFAAPLMHGGHSIRSDKENAGKDCLKNLEGCETCKLHFESEKQKENEKYLALESIVA SINEIDKASILSDTINYPRQLESRVAELESCTGSTDYEARSRSYMGMVDRTSDNHGIKKPWINKRKARDIDEAELELDEV APKDGMPVDLKVCMKEKEILIEMRCPYREYMLLDILDEANKRQLDVLSVHSSTLDGIFTLTLKSKFRGAAPVSPEGMIKQ ALRKTVGKT* |
| CDS seq | >Potri.014G083900.2.v4.1 ATGGCTCCTGTTGTCCAAGGCCAGCAGGTGGTGCCAGATAACTTGAGGAAACAGCTCGCAGTTGCTGTAAGGAGTGTGCA GTGGAGTTACGCAGTTTTCTGGTCACAGTCAACTAGACAACAAGGGGTACTGGAATGGGGTGATGGGTACTACAATGGAG ACATCAAGACTAGGAAAGTTGAAGCTATGGAGCTTAAAGCTGATAAAATAGGCTTACAGAGAAGTGAGCAATTGAGAGAA CTTTACGAGTCTCTCCTTGAAGGTGAGACCGGCCTGCAGGCTACGAGGTCTTCTCCTGCATTGTCTCCAGAGGACCTGTC AGATGAAGAGTGGTATTACTTGGTTTGCATGTCCTTTGTTTTCAATCCTGGTGAAGGGTTGCCAGGAAGAGCATTAGCTA ATAAACAGCCTATCTGGTTGTGCAATGCTCAATATGCAGATAGCAAAGTATTCTCTCGCTCTTTACTCGCAAAGAGCGCA TCAATTCAGACTGTGGTGTGTTTTCCCTATCTTGAGGGAGTAATAGAGCTAGGTGTGACTGAGCTGGTCACAGAAGACCC CGGTCTCATTCAGCATATCAAAGCCTCCTTATTAGACTTCTCAAAGCCTGTTTGTTCTGATAAATCGTTCTCTGCTGCTC ATAACAAGGATGATGATAAAGACCCAATGTCTACCAGGATCAGCCATGAGATAGTTGATACATTGGCTTTGGAGAATCTG TACACCCCAACAGAAGATATAGAATCTGAGCAGGAGGGAATTAACTACTTACATGGAAATGTGTGTGAAGAATTCAACAG GAACTCTCCTGATGATTTCTCAAATGGCTATGAGCACAATCTGGTGACTGAAGACTCCTTCATGCTTGAAGATTTAAAGG AAGGAGCATCTCAAGTTCAAAGTTGGCATTCCATGGATGATGAATTCAGCGATGATGTTAGAGATTCTATGAATTCAAGT GATTGCATATCAGAGGTTTTTGTGAAGCAAGGAAAGGTTGTACCCTCTTCCAAGGGAAAAGACATATCCCACCTGCAATT GAAAGTACTTCAAGAAGGAAACCACACAAAATTGAGCTCCTTAGATCCTGGAGCTGATGATGACTTGCACTACAGAAGAA CTGCATTTGTTATTCTGAAAAGCTCAAGTCAGTTGATCGAAAATCCATGTTTCCAAAGTGGTGATTACAAATCCAGTTTT GTTGGTTGGAAGAAAGGAGCAGCTGATGGCTATAAGCCACGAATACAACAGAAGATGTTGAAAAAGATTTTATTTGCAGC ACCTTTGATGCATGGTGGTCACTCTATTAGGTCTGACAAGGAAAATGCAGGAAAAGATTGCCTCAAGAATTTGGAAGGCT GTGAAACATGCAAGTTGCATTTTGAATCAGAAAAACAAAAGGAGAATGAAAAATATCTAGCACTCGAGTCAATAGTTGCT TCTATTAATGAGATTGACAAAGCATCAATCCTCAGCGACACCATTAATTACCCGAGACAGCTTGAGTCAAGAGTAGCAGA ACTGGAATCTTGCACAGGCTCGACAGATTATGAGGCCAGATCCAGGAGTTACATGGGCATGGTAGACCGGACATCAGATA ATCACGGCATCAAGAAGCCTTGGATAAACAAGAGGAAGGCCCGTGACATTGATGAAGCTGAACTAGAGCTTGATGAGGTT GCTCCCAAGGATGGCATGCCCGTAGATTTGAAAGTATGCATGAAAGAGAAGGAGATTCTCATCGAGATGAGATGTCCTTA CAGGGAGTACATGTTGCTCGATATCTTGGATGAAGCCAACAAACGGCAGTTGGATGTGCTTTCGGTTCATTCATCCACTC TTGATGGCATTTTCACGTTGACCCTGAAATCTAAGTTTAGAGGGGCAGCGCCAGTTTCACCAGAAGGGATGATCAAACAA GCACTTCGGAAAACTGTCGGCAAGACTTGA |
Pt_14:5389241-5389255:+ does not have available information here.