
| Microexon ID | Ps_NC_039367.1:62984239-62984252:+ |
| Species | Papaver somniferum | Coordinates | NC_039367.1:62984239..62984252 |
| Microexon Cluster ID | MEP37 |
| Size | 14 |
| Phase | 1 |
| Pfam Domain Motif | Trigger_N |
| Structure of Microexon-tag (flanking exon, microexon, flanking exon sizes) | 46,14,48 |
| Microexon location in the Microexon-tag | 2 |
| Microexon-tag DNA Seq | GYTSSTRCWGCACMRCCARTTCCWGGHTTYCGMAGRSWRAAAGGAGGRAAAACAYCARAKRTACCMARARRCWTYCTWYTRSARATKMTTGGWSMAKMMMRAGTBAMS |
| Logo of Microexon-tag DNA Seq | 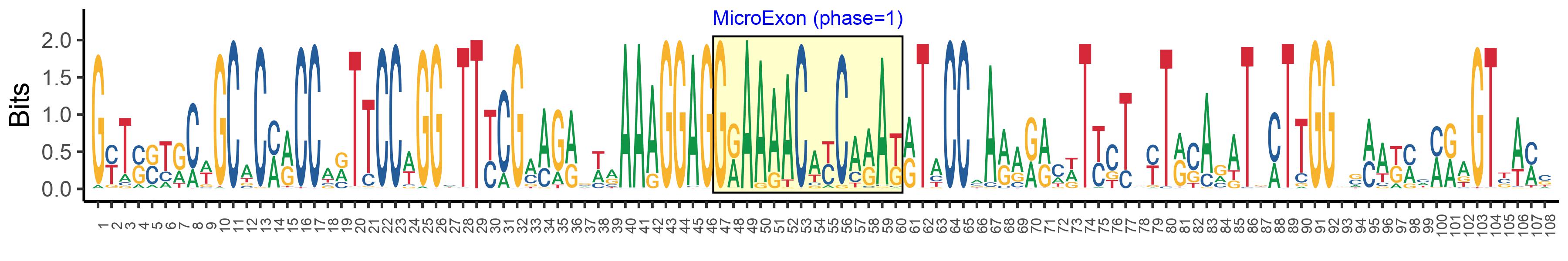 |
| Alignment of exons | 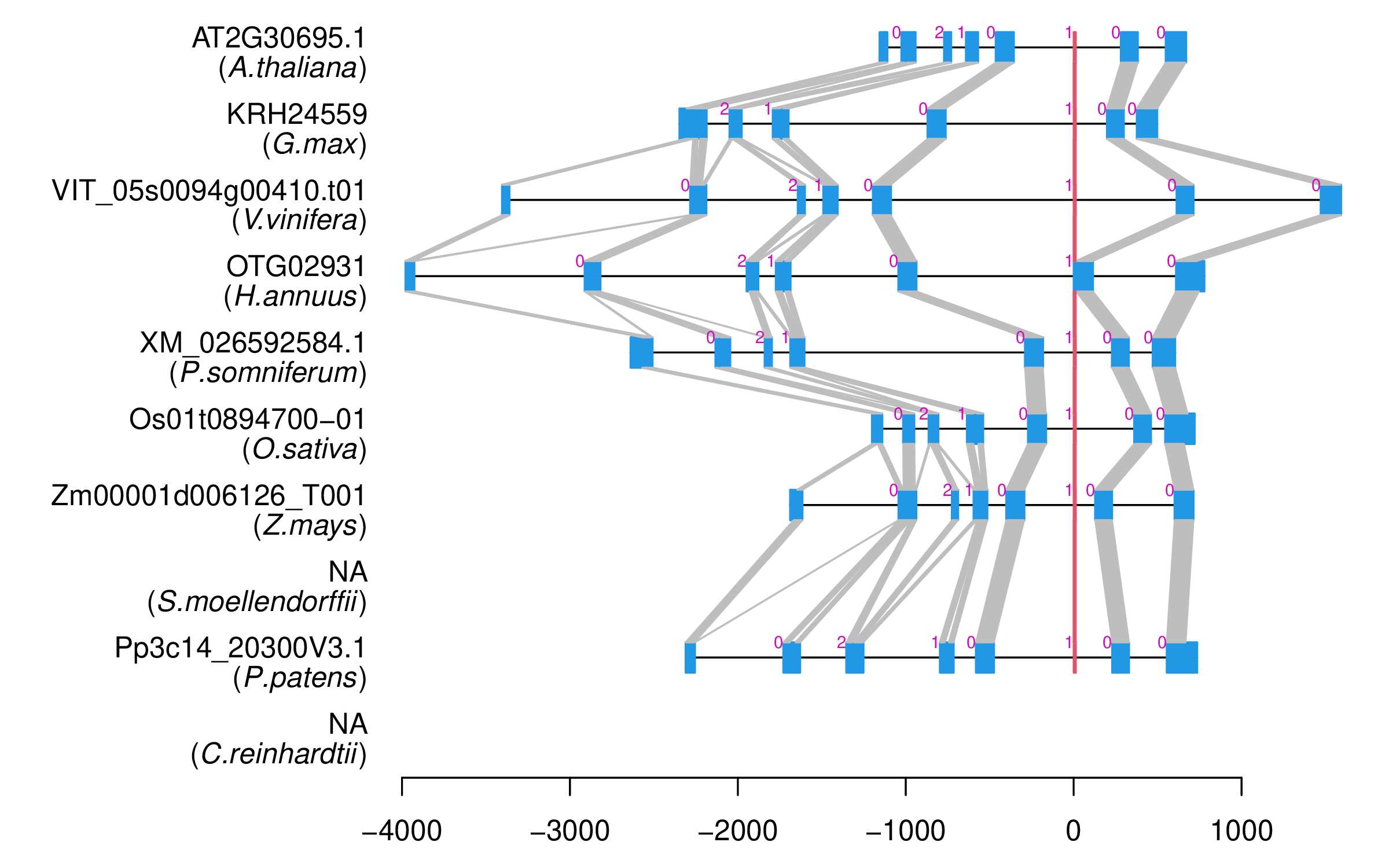 |
| Microexon DNA seq | GAAAGACATCCAAT |
| Microexon Amino Acid seq | GKTSN |
| Microexon-tag DNA Seq | GCTAAATCAGCACCACCTGTTCCTGGGTTCCGCAAAATGAAAGGAGGAAAGACATCCAATATTCCAAAGAGCTTTATATTACGGGCTCTTGGTGAAGATCGTGTGACC |
| Microexon-tag Amino Acid Seq | AKSAPPVPGFRKMKGGKTSNIPKSFILRALGEDRVT |
| Microexon-tag spanning region | 62984055-62984377 |
| Microexon-tag prediction score | 0.925 |
| Overlapped with the annotated transcript (%) | 100 |
| New Transcript ID | XM_026566437.1x |
| Reference Transcript ID | XM_026566437.1 |
| Gene ID | NA |
| Gene Name | NA |
| Transcript ID | XM_026566437.1 |
| Protein ID | XP_026422222.1 |
| Gene ID | LOC113318290 |
| Gene Name | NA |
| Pfam domain motif | Trigger_N |
| Motif E-value | 2.2e-08 |
| Motif start | 93 |
| Motif end | 233 |
| Protein seq | >XP_026422222.1 MACVSMSTVPSKTLSFPSIREVSIGTFNNPGHICGSTKSFSVHYTRNHNFPRLLFSSGESHRRSSQITYAVDSGVESSIT ADQDIVRHLKQAKVVIESRDDDKIQVRVDLTGIETLKIHDVVIANLAKSAPPVPGFRKMKGGKTSNIPKSFILRALGEDR VTKFTVQEIVSATMTEFVKTENLTVNNTFKTIQSAAELKATFKPGKEFGFNATLDIEKENSDQTSSSPSEVEVLPDTVKE NSDQTTSSPSEVEVLPQ* |
| CDS seq | >XM_026566437.1 ATGGCGTGTGTATCGATGTCTACAGTTCCTTCAAAAACCCTTTCATTTCCAAGTATTAGAGAGGTTTCTATAGGGACCTT TAACAATCCTGGGCATATATGTGGAAGCACTAAAAGCTTCAGTGTTCACTACACAAGAAATCACAATTTTCCTAGGTTAT TATTCTCAAGCGGCGAAAGTCACAGGCGATCATCCCAGATCACATATGCAGTTGATTCAGGAGTTGAGTCGTCAATTACT GCTGATCAAGACATTGTTCGTCACTTAAAACAAGCAAAGGTAGTCATTGAGTCACGAGATGATGATAAGATCCAAGTTAG AGTGGACTTAACTGGAATAGAAACACTAAAAATACATGATGTGGTTATAGCAAATTTGGCTAAATCAGCACCACCTGTTC CTGGGTTCCGCAAAATGAAAGGAGGAAAGACATCCAATATTCCAAAGAGCTTTATATTACGGGCTCTTGGTGAAGATCGT GTGACCAAGTTCACTGTACAAGAAATTGTCAGTGCAACCATGACCGAATTTGTAAAGACGGAAAATTTAACTGTGAACAA CACATTCAAGACAATACAATCAGCAGCAGAGCTAAAAGCGACTTTTAAACCAGGAAAAGAATTTGGTTTCAATGCTACAC TGGATATTGAAAAAGAAAATAGTGATCAGACCAGTTCAAGCCCCTCGGAGGTTGAAGTTTTGCCTGATACTGTGAAAGAA AACAGTGACCAGACAACATCAAGCCCCTCAGAGGTTGAAGTTTTGCCGCAATGA |
| Microexon DNA seq | GAAAGACATCCAAT |
| Microexon Amino Acid seq | GKTSN |
| Microexon-tag DNA Seq | GCTAAATCAGCACCACCTGTTCCTGGGTTCCGCAAAATGAAAGGAGGAAAGACATCCAATATTCCAAAGAGCTTTATATTACGGGCTCTTGGTGAAGATCGTGTGACC |
| Microexon-tag Amino Acid seq | AKSAPPVPGFRKMKGGKTSNIPKSFILRALGEDRVT |
| Transcript ID | Ps.61190.1 |
| Gene ID | Ps.61190 |
| Gene Name | NA |
| Pfam domain motif | Trigger_N |
| Motif E-value | 2e-05 |
| Motif start | 93 |
| Motif end | 181 |
| Protein seq | >Ps.61190.1 MACVSMSTVPSKTLSFPSIREVSIGTFNNPGHICGSTKSFSVHYTRNHNFPRLLFSSGESHRRSSQITYAVDSGVESSIT ADQDIVRHLKQAKVVIESRDDDKIQVRVDLTGIETLKIHDVVIANLAKSAPPVPGFRKMKGGKTSNIPKSFILRALGEDR VTKFTVQEIVSATMTEFVKTASYQT* |
| CDS seq | >Ps.61190.1 ATGGCGTGTGTATCGATGTCTACAGTTCCTTCAAAAACCCTTTCATTTCCAAGTATTAGAGAGGTTTCTATAGGGACCTT TAACAATCCTGGGCATATATGTGGAAGCACTAAAAGCTTCAGTGTTCACTACACAAGAAATCACAATTTTCCTAGGTTAT TATTCTCAAGCGGCGAAAGTCACAGGCGATCATCCCAGATCACATATGCAGTTGATTCAGGAGTTGAGTCGTCAATTACT GCTGATCAAGACATTGTTCGTCACTTAAAACAAGCAAAGGTAGTCATTGAGTCACGAGATGATGATAAGATCCAAGTTAG AGTGGACTTAACTGGAATAGAAACACTAAAAATACATGATGTGGTTATAGCAAATTTGGCTAAATCAGCACCACCTGTTC CTGGGTTCCGCAAAATGAAAGGAGGAAAGACATCCAATATTCCAAAGAGCTTTATATTACGGGCTCTTGGTGAAGATCGT GTGACCAAGTTCACTGTACAAGAAATTGTCAGTGCAACCATGACCGAATTTGTAAAGACGGCAAGTTATCAAACTTAA |