
| Microexon ID | Ta_7D:4291588-4291596:- |
| Species | Triticum Aestivum | Coordinates | 7D:4291588..4291596 |
| Microexon Cluster ID | MEP22 |
| Size | 9 |
| Phase | 1 |
| Pfam Domain Motif | Glyco_hydro_32N |
| Structure of Microexon-tag (flanking exon, microexon, flanking exon sizes) | 49,9,50 |
| Microexon location in the Microexon-tag | 2 |
| Microexon-tag DNA Seq | YSKYAMMGRACYGSYTWYCAYTTYCARCCYSMCAARAAYTGGATGAAYGATCCYAAYGGTCCAATGTWYTACAAGGGATKGTACCAYYTSTTCTAYCARTACAAYCCV |
| Logo of Microexon-tag DNA Seq | 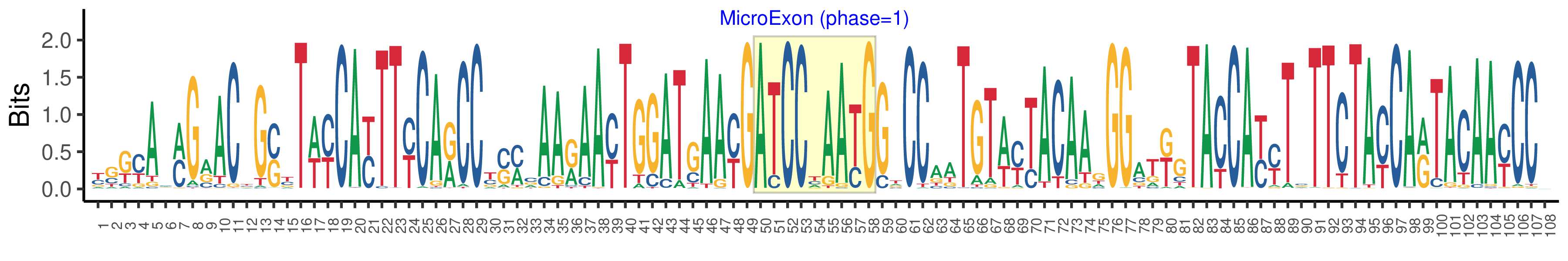 |
| Alignment of exons | 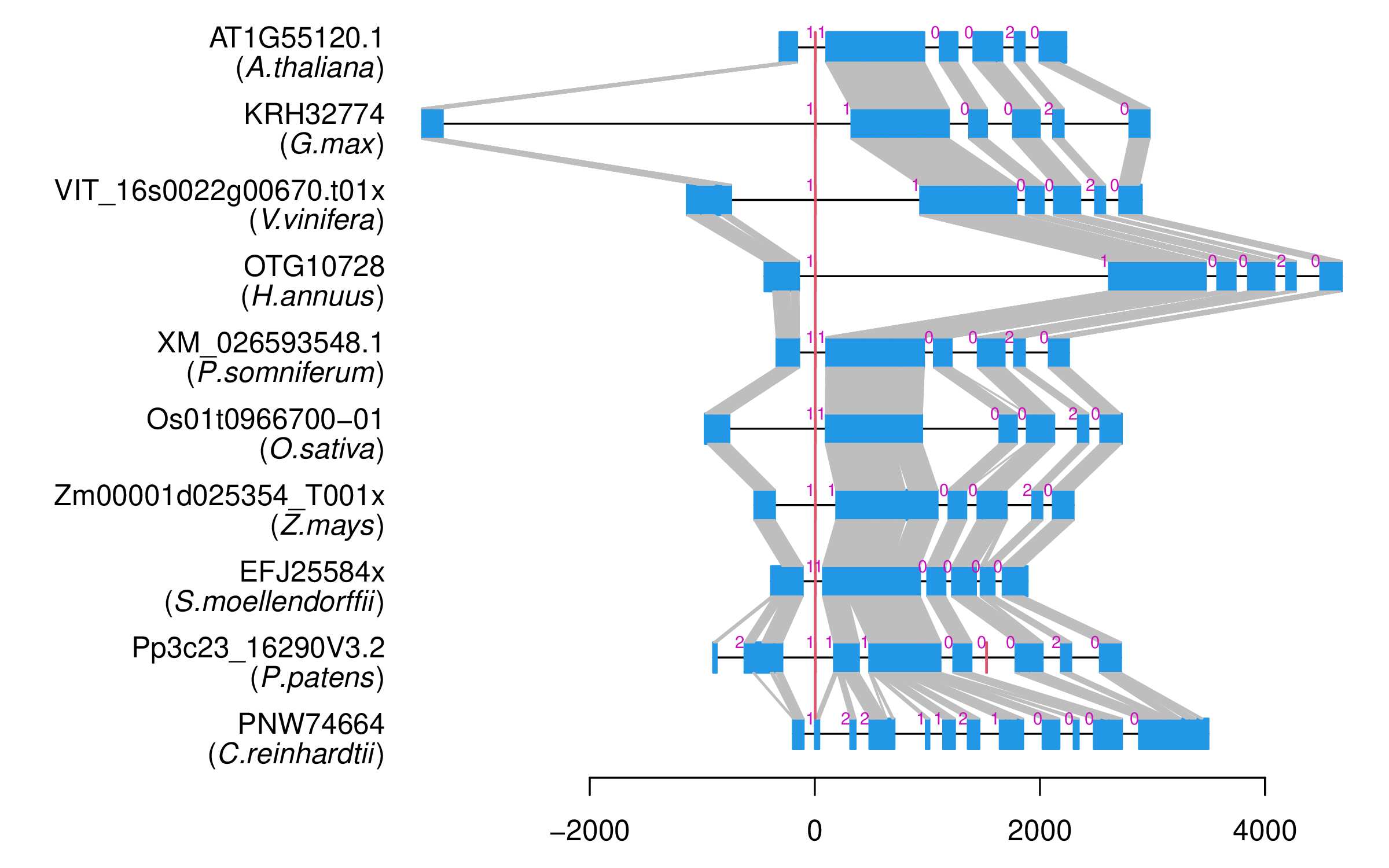 |
| Microexon DNA seq | ATCCCAACG |
| Microexon Amino Acid seq | DPNA |
| Microexon-tag DNA Seq | TGGCAGCGCACCGGGTTCCATTTCCAGCCCGAGAAGAACTACATGAACGATCCCAACGCTCCCATGTACTACCGTGGACGGTACCACTTCTTCTACCAGTACAACCCC |
| Microexon-tag Amino Acid Seq | WQRTGFHFQPEKNYMNDPNAPMYYRGRYHFFYQYNP |
| Microexon-tag spanning region | 4291419-4291789 |
| Microexon-tag prediction score | 0.9619 |
| Overlapped with the annotated transcript (%) | 100 |
| New Transcript ID | TraesCS7D02G008900.1x |
| Reference Transcript ID | TraesCS7D02G008900.1 |
| Gene ID | TraesCS7D02G008900 |
| Gene Name | NA |
| Transcript ID | TraesCS7D02G008900.1 |
| Protein ID | |
| Gene ID | TraesCS7D02G008900 |
| Gene Name | |
| Pfam domain motif | Glyco_hydro_32N |
| Motif E-value | 1.60E-97 |
| Motif start | 123 |
| Motif end | 437 |
| Protein seq | >TraesCS7D02G008900.1 MAPRDRRTTRSKDFLLSSGTATVAVKSPCAVPDTPPVPFAYGGLLYQQSGGRRRWRECTVVLGAVAMVALFVTHALLPRA SVLSEETPGETRAEQNIMVSAGADADGFPWSNEMLQWQRTGFHFQPEKNYMNDPNAPMYYRGRYHFFYQYNPTGVVWGNI TWGHAVSRDLVHWRHLPLAMVPDQWYDIHGVLTGSATILPNGTVIVLYTGKTDTSAQVQCLALPADPDDPLLVNWTKHPA NPVILPPPGIGLQDFRDPTTAWFDQSDLTWRTLIGSKDDNGHAGIALMYKTKDFIRYELIPRVLHRVEGTGMWECVDFYQ VGGGNNSSEEALYVLKASMDDERHDYYALGRYDAATNTWTPLDPELDVGIGLRYDWGKFFAATSFYDPVKRRRVMWAYVG ETDSLSANVAKGWASVQTIPRTVELDEKTRTNLLQWPVEEIETLRFNSTDFGVITIHNGSIIPLCLRQATQLDIEASFRL DTSASAIINEADVNYNCSTSGGASTMGVLGPFGLLIHATGNGGSEQLAVYFYVSRGRDGALRTHFCHDELLSSRANDVMK RVVGSTVPVLDGEALSVRVLVDHSIVESFAMGGRLTATSRVYPMEAIHMAAGVYLFNNATGSSITVEKLVVHEMASASYK QTFGAADA* |
| CDS seq | >TraesCS7D02G008900.1 ATGGCTCCTCGAGACCGCAGGACGACCCGATCAAAAGATTTCCTCCTGAGCTCCGGAACGGCGACCGTGGCCGTGAAGTC ACCCTGCGCCGTCCCGGACACGCCGCCGGTCCCGTTCGCGTACGGGGGGCTGCTGTACCAGCAGAGCGGCGGGCGGCGGA GGTGGCGCGAGTGCACCGTCGTGCTGGGCGCCGTGGCCATGGTGGCGCTCTTCGTCACCCACGCGCTTCTCCCAAGGGCC AGCGTGTTGTCCGAGGAGACGCCTGGGGAGACACGAGCGGAGCAGAACATCATGGTCTCCGCCGGCGCCGATGCCGACGG GTTCCCGTGGAGCAACGAGATGCTGCAGTGGCAGCGCACCGGGTTCCATTTCCAGCCCGAGAAGAACTACATGAACGATC CCAACGCTCCCATGTACTACCGTGGACGGTACCACTTCTTCTACCAGTACAACCCCACGGGCGTCGTCTGGGGCAACATC ACCTGGGGCCACGCGGTGTCGCGGGACCTGGTCCACTGGCGCCACCTTCCCCTCGCCATGGTGCCCGACCAGTGGTACGA CATCCACGGCGTCTTGACGGGCTCCGCCACCATACTCCCCAACGGCACGGTCATTGTGCTCTACACGGGGAAAACCGACA CCTCCGCTCAGGTCCAGTGCCTCGCCCTTCCCGCCGACCCAGATGACCCTCTCCTTGTCAACTGGACCAAGCATCCTGCC AACCCCGTCATCCTCCCGCCCCCCGGCATCGGCCTCCAAGACTTCCGCGACCCCACCACCGCCTGGTTTGACCAGTCTGA CCTGACATGGCGCACCCTCATCGGCTCCAAGGACGACAACGGCCATGCCGGCATCGCCTTGATGTACAAAACCAAAGACT TCATCAGGTACGAGCTCATCCCGAGGGTGCTCCACCGCGTCGAGGGCACTGGCATGTGGGAGTGCGTCGACTTCTACCAG GTCGGCGGCGGCAACAACTCCTCTGAGGAGGCCCTGTACGTGTTGAAGGCGAGCATGGACGATGAACGGCATGACTACTA CGCGCTTGGGAGGTATGATGCGGCGACCAACACATGGACACCACTGGACCCAGAGCTGGACGTGGGGATCGGGCTGAGGT ATGACTGGGGAAAGTTTTTCGCTGCCACGTCTTTCTACGACCCGGTGAAACGGCGGCGCGTGATGTGGGCCTATGTCGGC GAGACCGACTCCTTGAGTGCCAACGTTGCCAAGGGATGGGCTTCCGTCCAGACGATTCCGAGGACGGTAGAGTTGGATGA GAAGACCCGGACGAACCTCCTCCAATGGCCGGTGGAGGAGATCGAGACCCTCCGCTTCAACAGCACCGACTTTGGCGTCA TCACTATCCACAATGGATCCATTATACCTCTCTGCCTCCGCCAAGCCACCCAGCTTGACATCGAGGCCTCCTTTCGCCTT GACACTTCCGCTAGTGCCATCATCAACGAGGCTGACGTCAACTACAACTGCAGCACCAGCGGCGGAGCCTCAACCATGGG TGTGCTTGGTCCCTTCGGCCTCCTCATCCACGCTACTGGTAATGGTGGCAGCGAACAACTCGCGGTTTACTTCTATGTAT CGAGGGGCCGTGACGGGGCACTCCGAACCCACTTCTGCCATGACGAGTTGTTGTCGTCTCGGGCGAATGATGTCATGAAG CGGGTGGTGGGCAGCACAGTGCCGGTGCTCGACGGCGAGGCTCTATCTGTGAGGGTGCTCGTGGACCATTCGATTGTGGA GAGCTTCGCAATGGGCGGGAGGTTGACGGCCACGTCGCGGGTGTACCCGATGGAAGCCATCCACATGGCAGCGGGGGTAT ACCTTTTCAACAACGCCACCGGCTCCTCCATCACCGTCGAGAAGCTTGTAGTTCATGAGATGGCCTCGGCATCTTACAAA CAGACTTTCGGGGCCGCCGACGCCTAG |
Ta_7D:4291588-4291596:- does not have available information here.