
| Microexon ID | Gm_19:45265080-45265088:+ |
| Species | Glycine max | Coordinates | 19:45265080..45265088 |
| Microexon Cluster ID | MEP22 |
| Size | 9 |
| Phase | 1 |
| Pfam Domain Motif | Glyco_hydro_32N |
| Structure of Microexon-tag (flanking exon, microexon, flanking exon sizes) | 49,9,50 |
| Microexon location in the Microexon-tag | 2 |
| Microexon-tag DNA Seq | YSKYAMMGRACYGSYTWYCAYTTYCARCCYSMCAARAAYTGGATGAAYGATCCYAAYGGTCCAATGTWYTACAAGGGATKGTACCAYYTSTTCTAYCARTACAAYCCV |
| Logo of Microexon-tag DNA Seq | 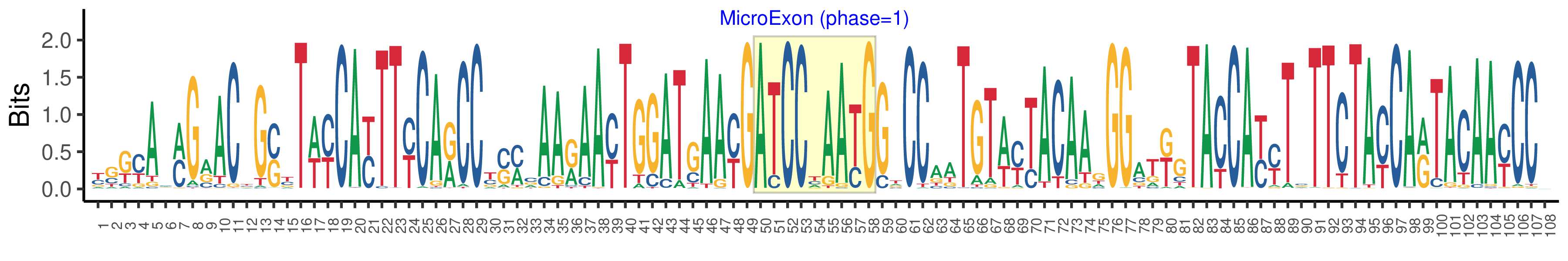 |
| Alignment of exons | 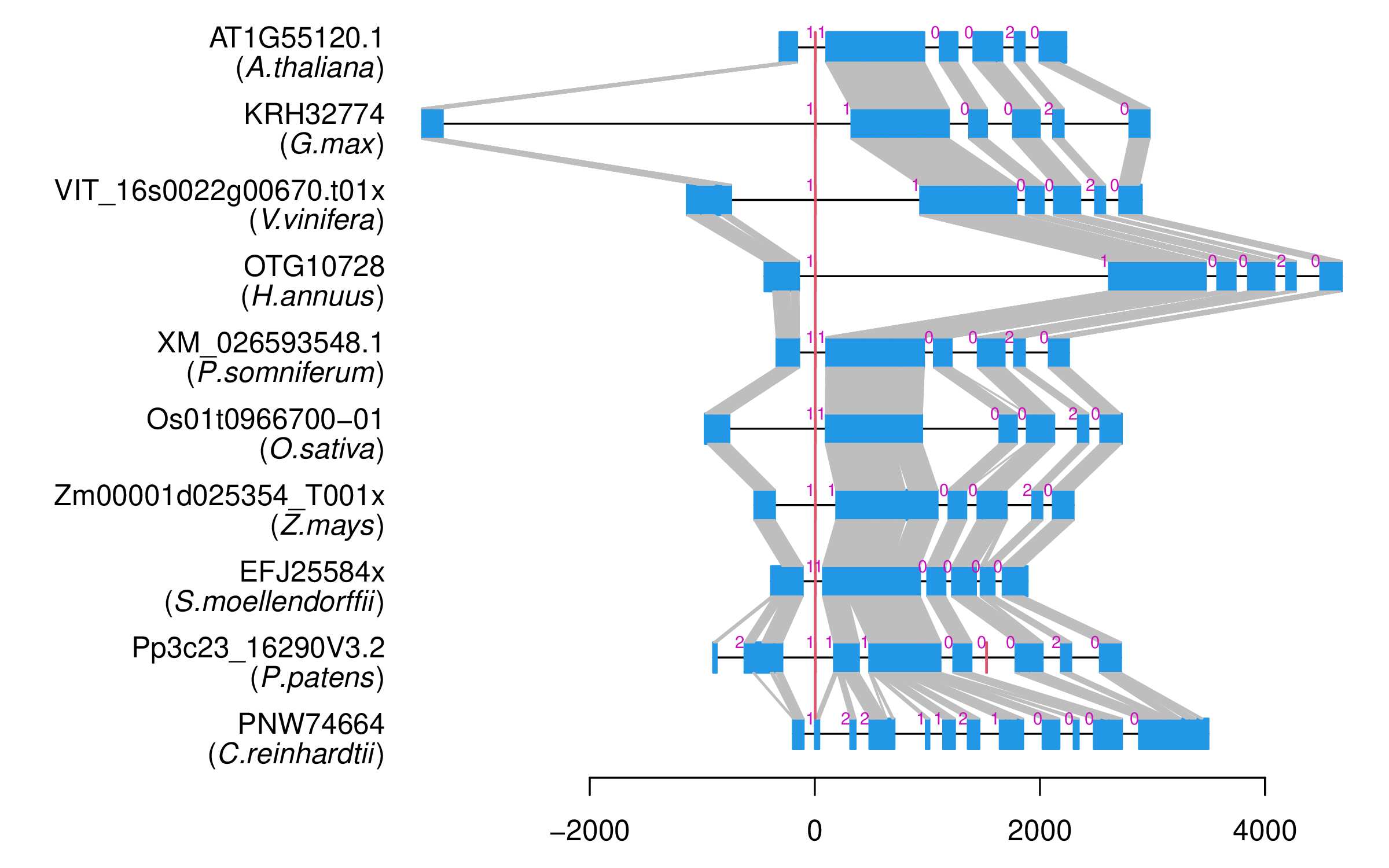 |
| Microexon DNA seq | ATCCAAATG |
| Microexon Amino Acid seq | DPNG |
| Microexon-tag DNA Seq | CAACACAGAACTGCATACCATTTCCAACCTCCTAAGAACTGGATCAACGATCCAAATGGACCCATGTACTACAATGGAATCTATCATTTATTCTACCAATACAACCCC |
| Microexon-tag Amino Acid Seq | QHRTAYHFQPPKNWINDPNGPMYYNGIYHLFYQYNP |
| Microexon-tag spanning region | 45263076-45265234 |
| Microexon-tag prediction score | 0.96 |
| Overlapped with the annotated transcript (%) | 45.37 |
| New Transcript ID | KRG96203x |
| Reference Transcript ID | KRG96203 |
| Gene ID | GLYMA_19G195400 |
| Gene Name | NA |
Gm_19:45265080-45265088:+ does not have available information here.
| Microexon DNA seq | ATCCAAATG |
| Microexon Amino Acid seq | DPNG |
| Microexon-tag DNA Seq | CAACACAGAACTGCATACCATTTCCAACCTCCTAAGAACTGGATCAACGATCCAAATGGACCCATGTACTACAATGGAATCTATCATTTATTCTACCAATACAACCCC |
| Microexon-tag Amino Acid seq | QHRTAYHFQPPKNWINDPNGPMYYNGIYHLFYQYNP |
| Transcript ID | Gm.28679.1 |
| Gene ID | Gm.28679 |
| Gene Name | NA |
| Pfam domain motif | Glyco_hydro_32N |
| Motif E-value | 5.6e-105 |
| Motif start | 52 |
| Motif end | 371 |
| Protein seq | >Gm.28679.1 MVLPKCRYITVVFFAFVVLLINNGVEAFHKVYPHLQSVSTISVSRQHRTAYHFQPPKNWINDPNGPMYYNGIYHLFYQYN PKGSVWGNIVWAHSISKDLINWRTLEPALYPSKPFDKFGCWSGSATIVPGKGPVILYTGVVDDKQTQVQCYAVPEDLNDP LLRKWVKPDKFNPILVANKGVNGSAFRDPTTAWWSKDGHWKILVGSRRKRRGIAYLYRSKDFMTWVQAKHPIHSKGETGM WECPDFYPVLVNGNQGLETSEGGNHVKHVFKNSLDMTRFDYYTVGTYFEDKDRYVPDNTSVDGWGGLRYDYGNFYASKSF FDPSKNRRILWGWANESDTKEDDVRKGWAGIQAIPRTVWLDSTGRQLVQWPVEELNNLRGKEVNMNSQKLQKGDYVEVKG ITAAQADVEVTFSFASLDKAETYDPKWVNAQDLCAQKGSKLQGGVGPFGLLTLASQNLEEFTPVFFRIFKGPVKHVVLLC SDATSSSLKSNMYKPSFAGFVDVDLATNKKLSLRSLIDHSVVESFGEGGKTNILSRVYPQLAVANQGHLFVFNNGTEPIS VENLKAWSMKPADIK* |
| CDS seq | >Gm.28679.1 ATGGTTCTTCCAAAGTGTCGCTATATCACTGTAGTGTTTTTTGCCTTTGTTGTGTTACTGATCAACAATGGCGTTGAAGC TTTTCATAAAGTATATCCTCATCTTCAATCTGTTTCAACGATTTCCGTGAGCAGACAACACAGAACTGCATACCATTTCC AACCTCCTAAGAACTGGATCAACGATCCAAATGGACCCATGTACTACAATGGAATCTATCATTTATTCTACCAATACAAC CCCAAAGGCTCAGTGTGGGGTAACATTGTGTGGGCCCACTCAATATCAAAGGATCTCATCAATTGGAGGACCCTTGAACC TGCCCTATATCCATCCAAACCATTTGACAAGTTTGGTTGTTGGTCTGGGTCAGCCACCATAGTCCCAGGTAAAGGACCTG TGATCCTCTACACCGGAGTTGTTGACGACAAACAAACTCAGGTTCAATGCTACGCCGTACCCGAAGACCTAAATGACCCA CTCCTCCGAAAATGGGTTAAACCTGACAAATTCAACCCGATCTTGGTTGCTAACAAAGGTGTTAATGGTAGTGCCTTTCG CGACCCAACCACGGCATGGTGGAGCAAGGATGGACACTGGAAGATATTGGTGGGTAGTAGAAGGAAGCGTAGAGGTATAG CTTATTTGTACAGAAGCAAGGACTTTATGACTTGGGTTCAAGCCAAACATCCGATCCATTCCAAGGGTGAGACAGGTATG TGGGAGTGTCCTGATTTTTATCCAGTTTTGGTTAATGGCAATCAAGGATTGGAGACGTCCGAGGGAGGGAATCATGTGAA GCACGTGTTTAAGAATAGTCTTGACATGACTAGGTTCGACTACTACACCGTAGGAACATATTTTGAAGATAAGGATAGAT ATGTCCCAGATAACACTTCAGTGGATGGTTGGGGTGGACTTAGGTATGACTATGGTAATTTCTATGCTTCAAAATCGTTT TTTGACCCCAGTAAGAACAGAAGGATCTTGTGGGGTTGGGCAAACGAGTCTGATACCAAGGAAGATGATGTTCGCAAAGG ATGGGCAGGAATTCAGGCAATTCCGCGAACTGTGTGGCTCGATTCTACTGGGAGACAATTGGTGCAATGGCCTGTTGAAG AATTAAATAATCTCAGAGGGAAAGAAGTTAATATGAACAGTCAAAAGCTTCAAAAGGGAGATTACGTTGAAGTAAAAGGA ATCACCGCTGCCCAGGCAGATGTGGAAGTTACATTCTCATTTGCAAGCTTGGACAAGGCAGAGACATATGATCCTAAGTG GGTAAATGCACAGGACCTCTGTGCCCAAAAGGGTTCAAAACTTCAAGGTGGGGTTGGACCATTTGGGCTTCTCACATTAG CTTCTCAAAATTTGGAGGAGTTCACTCCTGTGTTTTTTAGAATTTTCAAAGGTCCAGTTAAGCATGTGGTTCTCTTATGC TCCGATGCAACAAGTTCCTCTTTGAAGAGTAATATGTACAAGCCATCATTTGCTGGCTTTGTAGATGTGGATTTGGCCAC TAATAAAAAACTCTCTCTTAGGAGTTTGATTGATCACTCAGTAGTGGAGAGTTTTGGTGAAGGAGGGAAGACAAACATTT TGTCTCGGGTTTATCCACAGTTAGCAGTGGCTAATCAAGGTCACTTGTTTGTGTTCAACAACGGAACTGAACCCATCTCG GTGGAAAACCTCAAAGCATGGAGCATGAAGCCGGCTGATATAAAATAA |