
| Microexon ID | Gm_19:43773209-43773214:+ |
| Species | Glycine max | Coordinates | 19:43773209..43773214 |
| Microexon Cluster ID | MEP10 |
| Size | 6 |
| Phase | 0 |
| Pfam Domain Motif | DUF4788 |
| Structure of Microexon-tag (flanking exon, microexon, flanking exon sizes) | 51,6,51 |
| Microexon location in the Microexon-tag | 2 |
| Microexon-tag DNA Seq | TTTGARGAYTAYATYGADCCMCTYAAGRTKTACCTGRMTAGRTACAGAGAGWTGGAGGGTGAYACYAAGGGATCTGCWARRGSTGGWGATGSATCTGCTAARARRGAT |
| Logo of Microexon-tag DNA Seq | 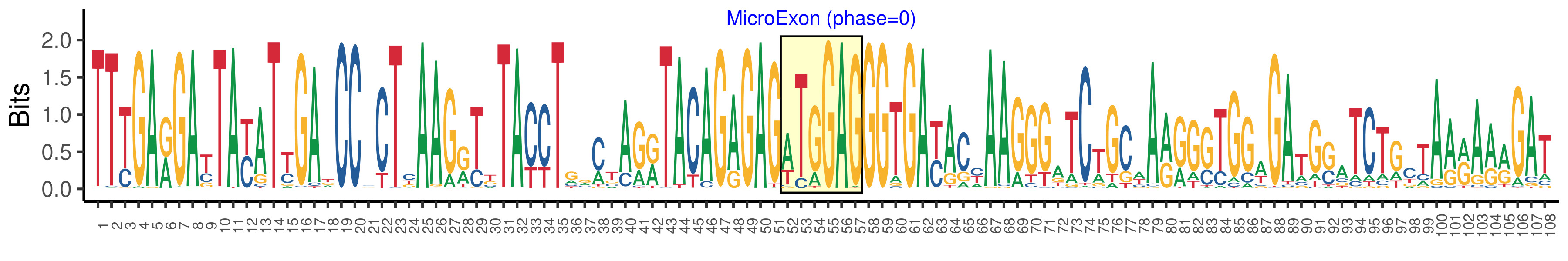 |
| Alignment of exons | 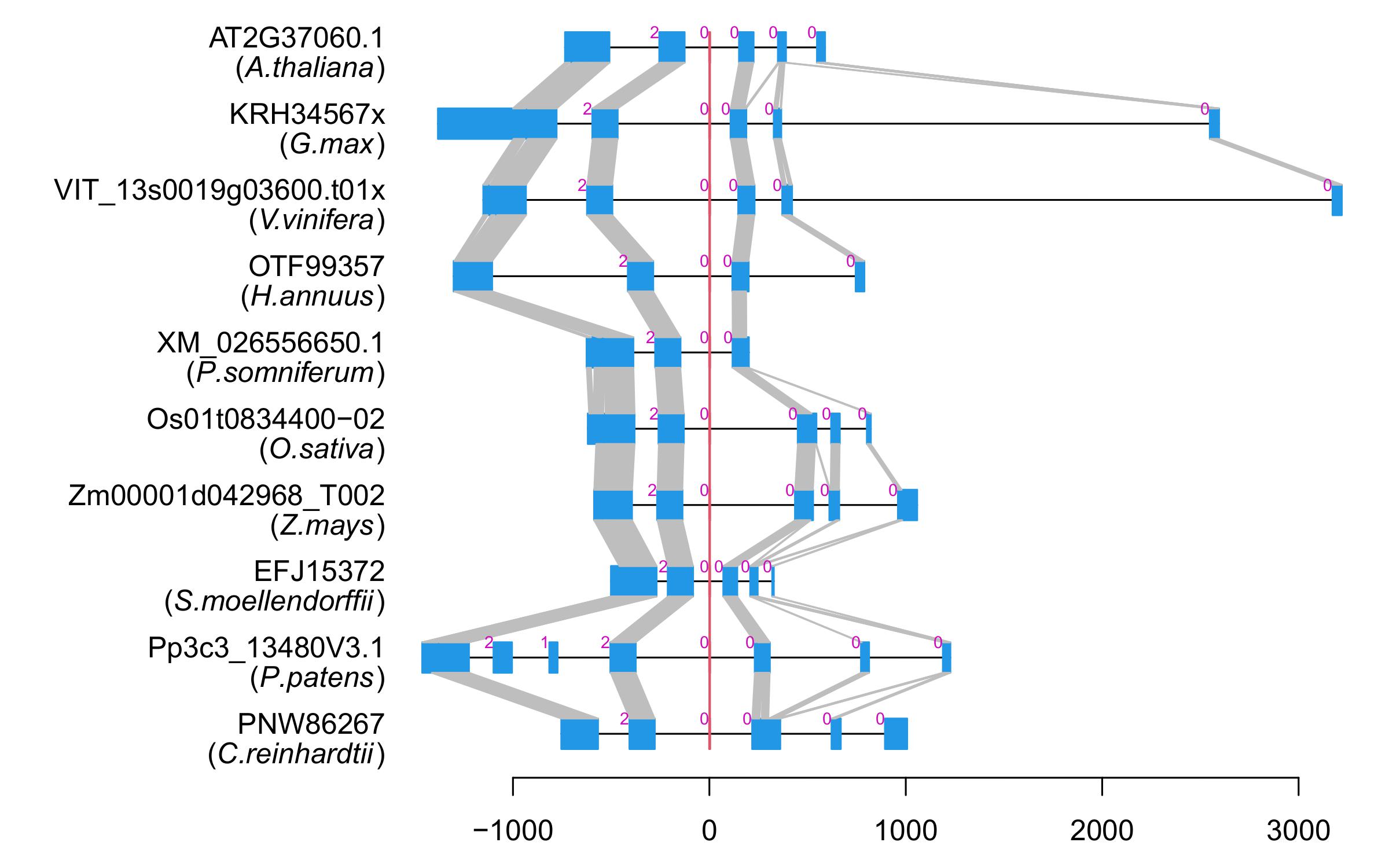 |
| Microexon DNA seq | ATGGAG |
| Microexon Amino Acid seq | ME |
| Microexon-tag DNA Seq | TTTGAGGATTATATCGATCCTCTTAAAATTTACCTCACTAGATACAGAGAGATGGAGGGTGATACGAAGGGTTCAGCCAAGGGCGGAGACTCATCTTCTAAGAAAGAT |
| Microexon-tag Amino Acid Seq | FEDYIDPLKIYLTRYREMEGDTKGSAKGGDSSSKKD |
| Microexon-tag spanning region | 43772965-43773378 |
| Microexon-tag prediction score | 0.9577 |
| Overlapped with the annotated transcript (%) | 94.44 |
| New Transcript ID | KRG95922x |
| Reference Transcript ID | KRG95922 |
| Gene ID | GLYMA_19G178500 |
| Gene Name | NA |
Gm_19:43773209-43773214:+ does not have available information here.
| Microexon DNA seq | ATGGAG |
| Microexon Amino Acid seq | ME |
| Microexon-tag DNA Seq | TTTGAGGATTATATCGATCCTCTTAAAATTTACCTCACTAGATACAGAGAGATGGAGGGTGATACGAAGGGTTCAGCCAAGGGCGGAGACTCATCTTCTAAGAAAGAT |
| Microexon-tag Amino Acid seq | FEDYIDPLKIYLTRYREMEGDTKGSAKGGDSSSKKD |
| Transcript ID | Gm.28496.1 |
| Gene ID | Gm.28496 |
| Gene Name | NA |
| Pfam domain motif | Unknown |
| Motif E-value | NA |
| Motif start | NA |
| Motif end | NA |
| Protein seq | >Gm.28496.1 MADGPASPGGGSHESGEHSPRSNVREQDRYLPIANISRIMKKALPANGKIAKDAKETVQECVSEFISFITSEASDKCQRE KRKTINGDDLLWAMATLGFEDYIDPLKIYLTRYREMEGDTKGSAKGGDSSSKKDVQPSPNAQLAHQGSFSQGVSYTISQG QHMMVPMQGPE* |
| CDS seq | >Gm.28496.1 ATGGCCGACGGTCCGGCGAGTCCAGGCGGCGGTAGCCACGAGAGCGGCGAGCACAGCCCTCGCTCTAACGTGCGCGAGCA GGACAGGTACCTCCCCATCGCTAACATAAGCCGCATCATGAAGAAGGCACTACCTGCGAACGGTAAAATCGCCAAGGACG CCAAAGAGACCGTTCAGGAATGCGTATCCGAGTTCATCAGTTTCATCACCAGCGAGGCCTCTGATAAGTGTCAGAGGGAA AAGAGAAAGACTATTAACGGTGATGATTTGCTCTGGGCCATGGCCACTCTTGGTTTTGAGGATTATATCGATCCTCTTAA AATTTACCTCACTAGATACAGAGAGATGGAGGGTGATACGAAGGGTTCAGCCAAGGGCGGAGACTCATCTTCTAAGAAAG ATGTTCAGCCAAGTCCTAATGCTCAGCTTGCTCATCAAGGTTCTTTCTCACAAGGTGTTAGTTACACAATTTCTCAGGGT CAACATATGATGGTTCCAATGCAAGGCCCGGAGTAG |